ACMG Annual Clinical Genetics Meeting 2022
Nashville, TN | March 22-26
Genomenon will be in Nashville for ACMG 2022, and we encourage you to connect with our team at booth 705 for a live demonstration of the Mastermind Genomic Search Engine. Also, register for our Exhibit Theater Presentation with Inozyme Pharma to see how Mastermind is connecting patients to approved therapies at the link below.

EXHIBIT THEATER PRESENTATION
Curating the Genome to Drive Precision Diagnosis in Clinical Care for Rare Disease
Speakers from Inozyme Pharma and Genomenon will discuss how to surmount obstacles related to translating sequencing data into actionable results, and present Genomenon’s comprehensive curated variant content as the approach for accelerating rare disease diagnostic workflows and connecting patients to clinical studies and approved therapies.
EXHIBIT THEATER SPEAKERS

Catherine Nester
Vice President, Physician and Patient Strategies, Inozyme Pharma

Dr. Mark Kiel
Founder, Chief Scientific Officer, Genomenon
ACMG RESOURCES
We love free stuff too.
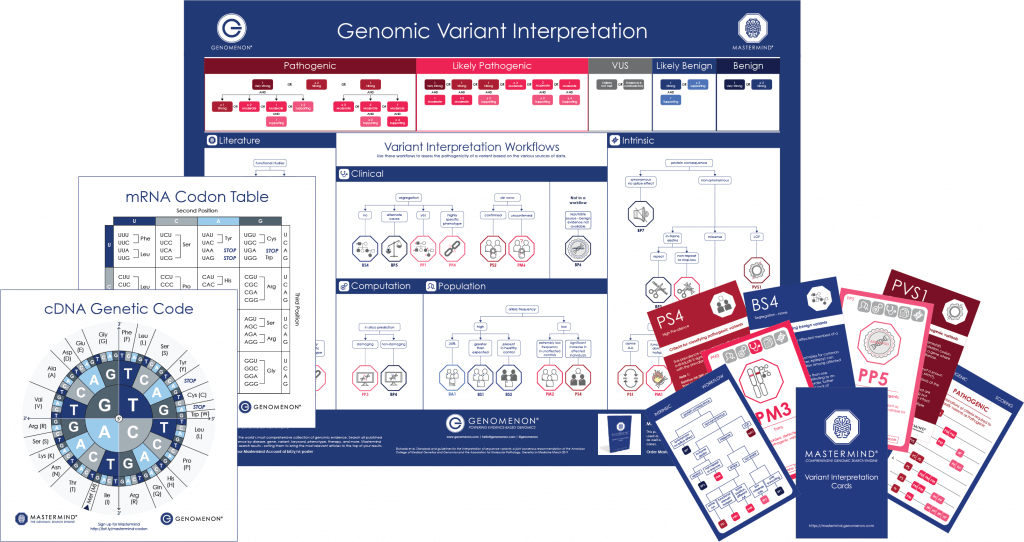
Complete the form to download your variant interpretation resources. Decorate your workspace with our ACMG Poster, or test your knowledge with our Variant Interpretation Playing Cards. Even better – get your own copy of our popular cDNA and mRNA Codon Charts.
Meet us at Booth 705
Connect with members of our leadership team at ACMG 2022! Talk to a Mastermind Specialist and:
Demo the world’s most comprehensive source of genomic evidence
Schedule 1:1 meetings – on-site or virtually
Learn about extended trials and special offers for clinical labs
Learn about our comprehensive genomic landscapes for Pharma

Create your FREE Mastermind account and start searching!
GET MASTERMINDWEBINAR TRANSCRIPT
MARK: Well, thank you, everybody, for attending, physically or virtually! Today we will be learning about how curating the genome can inform and improve genetic diagnosis for rare disease. My name is Mark Kiel, and I’m the Chief Science Officer and a founder of Genomenon. I’m very privileged to be here speaking with Catherine Nester, a friend from Inozyme, who is the VP of patient and physician strategies there. She and I will be exemplifying how curating the genome can help diagnose rare disease patients, with a specific use case for ENPP1 deficiency.
My role here is to introduce Genomenon and our clinical software Mastermind, and then let Catherine take it away to go through the clinical and diagnostic features of this disease. So, a little bit about Genomenon — Genomenon is a genomics intelligence company, and our mission is to save and improve lives by making genomic information actionable. Our focus is on ensuring that there is comprehensive and accurate evidence for our clinical and research users, to ensure that they have accurate understanding of these genetic diseases, both diagnostically and from a biochemical understanding. Clinically, the way that we do this is through a product called Mastermind. Mastermind is a genomic search engine that allows our clinical users to quickly identify references for patient diagnosis and for treatment decisions. This evidence comes from a comprehensive collection of clinical and functional references, including eight million full text and three million supplemental data sets, and comprises nearly 15 million variants.
This happens through a core technology that we refer to as genomic language processing. I call this the “order from chaos” slide. On the left is the chaos of the scientific and clinical and medical literature, those many millions of references and data sets that I alluded to earlier. The order that we bring to this chaos is through genomic language processing, recognizing those entities that are necessary for understanding what this all means: diseases, phenotypes, therapies, genes and their genetic variants, as well as categorical key terms for clinical and functional studies, no matter how an author may describe any of those, including the complicated genetic nomenclature for recognizing variants. We put all that index data together into what we call genomic associations to know how those diseases phenotypes and therapies interrelate with those genes and variants.
So that’s one of our core technologies. That falls under the rubric of artificial intelligence or computational intelligence. That’s on the left, that is to ensure the maximal sensitivity for understanding this information, so genomic language processing and those genomic associations which are automatically indexed. Another side to our capability is exemplified there on the right, and that is with our expert team of curators, powered by an AI-driven automation curation capability, we can review all of that evidence and ensure the utmost accuracy and specificity of that information, to have a unique combinatorial balance approach, to ensure both maximal sensitivity and specificity and applicability of that information. Clinically, what that amounts to, from the perspective of curating this information, is what we refer to as disease-specific curated content, which, for a select and growing number of genes, is now available in the Mastermind software to inform clinical diagnoses. What this means, as we’ll talk through for this gene and this disease in particular, is every published ENPP1 variant, every relevant reference publishing any one of those variants, clinical cases, cohorts, functional studies, as well as any relevant database information for each of those variants necessary and sufficient to inform a clinical interpretation of those curated data sets.
This is what that looks like for one of those genes. You see how there’s a clear takeaway summary of that curated information. I call this the “action from order” slide, so we go from chaos to order to action. This is one of those genes, and I’ll invite Catherine Nester to the stage to speak about the ENPP1 gene specifically. Catherine?
CATHERINE: Thanks for having us! When we started our partnership with Genomenon, we really were looking for a way that we could get a faster diagnosis for patients, and then also, how could we connect the patient and the physician to a clinical trial? That was sort of the onus for how this work got kicked off together. I’m going to talk a little bit about ENPP1 deficiency. What is this disease? Some of you in the audience may know this as either GACI or ARHR2. It’s a disease that really is across the spectrum. There’s a lot of heterogeneity to the disease. The hallmarks of it are ectopic calcification of the arteries, organs, joints, and ligaments; pathologic skeletal mineralization; neointimal proliferation; and vascular stenosis.
In a healthy population, what does the role of ENPP1 really play? It plays this critical role in regulating mineralization and neointimal hyperplasia. How that happens is that ATP is transported intracellularly to extracellularly. ENPP1 then breaks that down into pyrophosphate (PPi) and AMP. Then, it’s further hydrolyzed by CD73 into adenosine and phosphate. For a healthy population, that really is what controls the ectopic growth and formulation of hydroxyapatite. It inhibits the mineralization, maintains healthy bones and teeth. Then, from a vessel perspective, it inhibits neointimal hyperplasia. You see where this could be going if this isn’t working the right way. For patients who have ENPP1 deficiency, they have a bi-allelic autosomal recessive disease, and so they don’t have this enzyme activity. The result of that is this low level of inorganic pyrophosphate. It’s really the best biomarker of the disease, although right now, it’s not that easy to measure. Really, what these low levels of PPi leads to is mineral deposits in arteries, joints, and organs. Patients will then go on to have osteomalacia, rickets, reduced bone strength, and then these narrowed artery lumens as well.
I mentioned the heterogeneity of it in infancy. This is the most mortal time of the disease. For infants, about 50% of them will die in the first six months of life. It’s usually from these severe cardiovascular complications. This is the phenotype known as GACI, or Generalized Arterial Calcification of Infancy. For the 50% of these babies who survive, this disease then starts to evolve. In kids and adolescents, we really start to see the skeletal phenotype emerge. They have rickets, they have hypophosphatemia,
they start to have these calcifications of their joints and ligaments, bone deformity, hearing loss. Then, in adulthood, it’s really about osteomalacia. The hypophosphatemia continues, and the adult population really has bone and joint pain and some impaired mobility. We’ve been contacted by some patients who did not get the correct diagnosis until the third, fourth, and fifth decade of life, and so they lived with these symptoms for a really long time, but knew no real cause for it.
I talked a little bit about the mortality. This is from Dr. Rutsch and his team. He’s sort of one of the godfathers of all of this work. He looked at a natural history study of these 55 patients, and in this particular cohort, 65% of these babies died, either in utero, or within the first six months of life, so again, very mortal. You can see on the right there, the predominant calcification was aorta, coronary arteries, pulmonary arteries, etc. If you’re looking in the prenatal period, ideally, we would find these babies prenatally, and we often see fetal distress or polyhydramnios. They’ve got effusions and hydrops.
I mentioned, for those who survived that critical period, this is again from Dr. Rutsch and his team, we had serial measurements for some of these kids that survived. What we were looking at were their serum phosphate levels, and then maximum renal tubular phosphate reabsorption. As you can see, all eight of these children had hypophosphatemia and hyperphosphaturia. You see how this disease is evolving, you see where this is gonna to be heading. One thought here is, why does this become then a skeletal disease? Well, FGF23 starts to rise in these babies, and the thought is that that is is a compensatory mechanism for the calcification that’s happening. We see these high FGF23 levels in this population as they grow and develop. This is just another cohort of patients for which we had some some serial measurements. What then happens, when all of this is going on, we see these hypophosphatemic rickets. This is actually looking at a larger cohort, 84 patients from Drs. Rutsch and Ferreira. You see on the left, a seven-year-old with genu valgum. You see in the middle there, a 49 year old who’s got bowing of both femurs. Then on the right, you see the metaphyseal fraying and flaring in the knee. This is really just, you know, highlighting how the disease continues to evolve over time, and becomes sort of this skeletal phenotype.
So how do we differentiate it? Well, it’s not that easy. All of you sitting in the room, the genetics community, plays a huge role in getting the right diagnosis for these patients. If you look at them and you compare them biochemically to a patient who has, for instance, XLH, biochemically, they are going to look pretty much identical to each other. Biochemical profiling isn’t really going to be that helpful for trying to figure out which disease this might be. The clinical features, there are some places where the clinical features really overlap, such as the rickets and osteomalacia, the short stature, gait abnormalities, the bone and joint pain. Both populations will have enthesopathies and some hearing loss. The difference in ENPP1 deficiency is the enthesopathies and hearing loss generally occur a little earlier than perhaps in XLH. Then, of course, on the right, the differentiating characteristics are the arterial, organ, and joint calcification, and those other things listed there. Some other clinical features: Calcification and fusion of the cervical spine ligament is pretty common in this disease, hyper-cementum and delayed shedding of teeth — different than some some other diseases that you may know, like HPP. These kids retain their teeth, and sometimes when they’re having orthodontia work, it doesn’t go so well, because they’ve got this hyper-cementum. Then, of course, the arterial calcification and the cardiovascular complications.
So the diagnosis takes a multi-disciplinary team. Good history, good family history. Looking at all of those clinical features to try to find the things that are unique to ENPP1 deficiency, I mentioned the laboratory values that are important here. But at the end of the day, it’s genetics, right? We need to make sure that all of these patients for which you have a high suspicion get sequencing. There were some diagnostic pathways that have been developed with some input from some of the expert groups, who would see these kids across the age spectrum.
Here’s the pathway for in utero and for the infant population. I’m happy to provide these slides to whomever in the room would like to have them. That’s perfectly fine to share that with you. The pathways really are focused on the physician group that may see the patient at that point. Then, also, the presenting symptoms that are common to the patient at that time in their life. Here is the one for the kids, and you’ll see, the infant one is really about the cardiovascular issues of the disease. The adolescent pathway is more about the skeletal. We sort of lead with the skeletal component of the disease, because that’s most likely how they’re going to get picked up in that age group. If you can get inorganic pyrophosphate measurements at your institution, that’s great, that is helpful for the disease. It is the best biomarker for the disease, but it’s not always that easy to get. Of course, as I mentioned before, it really is about the genetics. As you’ll see here, we also mentioned ABCC6 mutations. Generally with these kids, it’s driven by mutations in ENPP1. There is a small population called GACI type 2. Those kids look a lot like the GACI type 1 babies, but that mutation is actually in ABCC6. Then, here’s the pathway for the adult population, and again, as you see, it’s really driven by the skeletal features of the disease.
So why are we talking about this, what’s the importance here? We are developing an enzyme replacement therapy for the disease. It is in clinical studies already in an adult population. This is a recombinant human ENPP1. The construct is a recombinant Fc fusion protein with a soluble extracellular domain. Right now, the dosing is twice a week subcutaneously. What we’re going to do is an ascending study, so each cohort is going to get a higher dose, and this whole study in adults is about of the safety and the dosing, PK/PD, all of that. The hope is, once we are finished with this study, we’re going to go into infants and children as quickly as possible. Of course, that’s really where the urgent medical need is. With that, I’m gonna turn it back over to Mark.
MARK: Thank you so much, Catherine! I’ll bring this story together here. For those of you who joined late, we’re again talking about Genomenon’s ambition to curate the entire genome, gene by gene and disease by disease, with the example here being ENPP1 deficiency and GACI. This is a schematic that I’ve put together to illustrate the power of comprehensive curation of this evidence and expert clinical-grade interpretation of all these genetic variants, across all of ENPP1 and ABCC6, and then, every rare disease and cancer-associated gene. Just to walk you through this schematic, each of those bars in the bell distribution are genetic variants with their different classifications: benign, likely benign, variant of uncertain significance, likely pathogenic and pathogenic. The benefit of comprehensively curating all of this reference material, clinical and functional studies, is twofold. You have a maximal understanding of all of the causative genetic variants, so all the bars rise, but in addition to that, you have the most comprehensive reflection of all of that evidence. Any of that clinical or functional evidence that comes out of comprehensively curating the genome will push the bars appropriately to the left, toward pathogenic, for all of the cases that contribute evidence to these interpretations for all the cohort studies and all of the functional studies. More variants, more evidence, means more patients more accurately diagnosed. As Catherine illustrated, that is a critical need, particularly for diseases like ENPP1 deficiency that are diagnostically challenging, where ultimately, the diagnosis culminates in a confirmation at the genetic level. That’s where Genomenon figures into this effort, in the service of helping identify and treat more patients.
So I mentioned — this is the action slide that I talked about earlier — below in blue is what you typically see in a Mastermind search for any one of these genetic variants. As we’ve been alluding to, Genomenon has curated all of these ENPP1 variants and curated all of the evidence associated with each of those, and now are presenting the fruits of that work for any clinician or laboratorian who performs a search in Mastermind. You see again, the banner at the top will illustrate, for any one of those ENPP1 variant searches, what the result was, what the final interpretation was, and a summary breakdown of what the evidence was that culminated in that call.
But we go further. This is the interpretation view of that evidence. I realize it’s small, but I invite you to perform a search on an ENPP1 variant in Mastermind to see this in action. What you’re seeing here is that summary at the top, enumeration, quantitation of the different categories of evidence, the numbers of cases, the numbers of functional studies in summary form there that comport with the ACMG clinical interpretation criteria. Then below that, you see each one of those references reference-cited within those categories, clinical, functional, case, cohort, etc., sentence context from each of those specific studies that justify the curation, and the resultant interpretation of each of those studies, and tags to associate those with the specific diseases that those patients had, or in the case of functional studies, what those functional outcomes were. In addition to all of the relevant database information, as I said, that’s a necessary component of this accurate interpretation. There’s additionally places for you to add your own comments to share with your colleagues, either internal to your institution or external.
The key takeaway from this is that now, we’re talking not just about organizing all this evidence, but having it pre-interpreted for a provisional call, so that we’re shortening the time from sequencing a patient to confirming the diagnosis. Furthermore, there’s a call to action here. There’s information that is provided to you for this select set of genes, ENPP1 included, where there’s notification of a trial. There are further links to additional information that help you understand the diagnostic challenges, connecting you with a network of individuals who can help support you in ensuring that this patient’s diagnostic and therapeutic journey is accurate and efficient.
So again, for those of you who joined late, we referred to this in Mastermind as disease-specific curated content for genes like ENPP1, diseases like GACI, and a growing number of additional genes and diseases. We’re talking about, now, every published variant in those genes, every pertinent reference expertly curated, and all of the relevant database information necessary to result in an ACMG clinical-grade interpretation of this information and its applicability for diagnostic and therapeutic decisions. There’s a link there that you can follow to have access to Mastermind for a trial period, or for the basic edition indefinitely. I exhort you, any of those of you who are interested in learning more about ENPP1, disease-specific curated content, the work that Inozyme is doing, or the work that Genomenon is doing — our booths are adjacent to each other over in the 700 block. Anybody who’s scanned here will receive a gift from Genomenon. We welcome traffic over to the Genomenon booth to discuss what we talked about here, as well as to the Inozyme booth to learn more about GACI.
With that, I’ll take any questions, or any questions about GACI in particular for Catherine. Yes.
RITA: Hi there, this is Rita Quintana representing the clinical genomics R&D team at Natera. I have a question about Genomenon. Well, maybe two questions: How many attributes have you curated of genes and diseases, and how many gene-disease pairs have you curated at the moment?
MARK: So this is a new initiative that we’ve undertaken. We’ve been curating this information for quite a while now, but releasing it into our clinical software is a relatively new endeavor. We’re very happy to be announcing the curated content for ENPP1 today, at present, it’s two dozen different genes and conditions. Again, we welcome you to join our teammates at the booth to learn more about which specific diseases those are, but we’re exponentially accelerating the curation and release of that content into our software.
RITA: Okay, thank you, and then, a third question: from my prior experience, because I now work mostly on R&D, not in production, but I have years of working in production as a genomic scientist interpreting cases. So I noticed that your UI is very busy.
MARK: Well, thank you.
RITA: I wonder, from my personal experience, because as a scientist, you need many pieces of information coming to you, sometimes sequentially and not at the same time. Like, the fonts, even in this bigger view, they’re kind of small. Maybe I don’t need all the pieces of information at the moment because how my brain is processing the information. Maybe I need three pieces of information, so I wonder if you have any plans to revamp your UI, based on user experiences?
MARK: Yeah, this is great feedback. I have a very large monitor at home when I work on this data, so I recognize that not everybody has that setup. I will call out, though, that there is a cascade of the information. I talked about the banner, the key takeaways, the final provisional call and the categories of evidence that culminated in that call. We very much take a trust-but-verify approach with our curated content, so you can track down to each of these references and examine that information at any depth that you need. That’s very much our philosophy behind presenting this information. We’ll take cues from you for that user input about the font size and the configuration.
RITA: Yeah, I mean, this is just one user, but maybe from your community of users, you can use that information to adjust.
MARK: Yes, I welcome it. That’s excellent feedback, thank you very much. As you say, especially as the information gets more and more dense, we will be able to accommodate user experience.
RITA: It’s just because I have used as many software options as possible, as are available, and I noticed like the main issue is: if you look at good applications like Asana or Ocean, you see the flow. They invest a lot of time in the UI, and that helps you sell the software in a way.
MARK: Great, thank you so much! Thank you. Please stop by, we’ll talk further — I’m mindful of the time. I think we’ve got plenty of time.
AUDIENCE: How frequently is the information updated?
MARK: Yeah, excellent question. The data content in Mastermind is updated on a weekly basis, and the curated content is updated on a yearly basis. Putting those two together, you’ll see the interpretation that is reviewed on a yearly basis, but you’ll see any additional information that has come in since that time on a weekly basis. As we scale this operation up, there’s interest in us accelerating the periodicity of those curated updates. But the core content is updated on a very regular weekly basis. Any other questions? Thank you, Catherine for doing that.
AUDIENCE: Hi, the question I have is — what I’m seeing is quite similar to what, for example, HGMD is presenting for many years already. It’s a little more phenotypic correlation, but so far, that’s pretty much the only difference I see. Is it worth the effort for something that’s well established already and comprehensive, unlike a couple of dozen genes that you currently have. What’s the added value? We are working on interpretation, now clinical interpretation of rare disease cases every day. That’s core business, and so far, we would really love to have something better than HGMD.
MARK: We got it for you. Well, let me address your questions specifically. This is the slide that I’d like to use to exemplify my answer. It is the value of the comprehensiveness of a computational approach. We talked about the periodicity of the updates. There’s so much information coming out so frequently about so many diseases that it defies staying up to date, even with an extremely large team of manual curators. To solve that challenge, we have built out that computational intelligence to ensure that every variant is found, so that you don’t leave any variants behind. That’s the maximal sensitivity that we talked about, and I challenge you to look at our sensitivity compared to HGMD’s.
Secondly, though, and very importantly, HGMD shows the first study, or maybe the first two, maybe a review article. What we’re talking about here,though, is not just a comprehensive reflection of all those variants, but also, of all the evidence. Even if it’s from last week, the new case where we talk about a functional study taking you from sub-clinical pathogenicity in a VUS, one case can provide strong evidence for pathogenicity. That will tip the balance for you to better understand whether this is a diagnostic gene. That keeps going on and on, week over week. We’re not stagnant like HGMD is, when they review that information and keep it at the first seminal reference. This is updated on a regular basis, and you’ll see the benefits of that when you do a lookup at specific variants. There’s a great deal of value in having a comprehensive reflection of that evidence.
CATHERINE: It actually changed the call on some of the variants that were found, and so we now have significantly more pathogenic or likely pathogenic variants than we had when the project started. For us, that was a huge value.
MARK: That is true of the variety of different genes that we have already curated: an increased number in variants, but also, an increased number of pathogenic variants as a result of that comprehensive evidence.
I think I’m getting the cane — we’re at booth 705 If you’d like to talk more. Thank you all very much for attending, and I welcome some more interaction at the booth. Thank you!