ACMG Annual Clinical Genetics Meeting 2024
Toronto, ON | March 12-16
Meet the Genomenon team in Toronto, Ontario for ACMG 2024, the premier medical genetics event.
We’re excited to meet with many of you at ACMG this March. Our scientific talks, posters and booth will showcase how we are simplifying complex genetic data into actionable insights for patient diagnosis and precision medicine development. Meet with us to learn how Genomenon can provide you and your teams with genomic intelligence through our software, services, and data.
SOFTWARE
As the world’s most comprehensive source of genomic evidence, the Mastermind Genomic Intelligence Platform enhances variant interpretation by reducing turnaround time, increasing diagnostic yield, and accelerating throughput.
SERVICES
In tandem, Genomenon’s Curate Pro and Interpret research and data services further empower variant interpretation teams, enabling them to operate with heightened efficiency through rapid, on-demand evidence curation and classification.
DATA
Genomenon’s tech-enabled team of scientists has curated variants across hundreds of genes and our API gives you access to all of our indexed variant data to help with your clinical workflow automation.
Join us for our Exhibit Theater presentation that will demonstrate how Genomenon is utilizing these tools and services to fully curate the human genome and provide critical insights for patient diagnosis and precision medicine development.
WATCH NOW | Genomenon’s Scientific Platform Presentation
and Exhibit Theater talks
EXHIBIT THEATER PRESENTATION
The Fully Curated Human Genome – Implications for Improved Clinical Diagnostics for both Known and Novel Variants
Diagnosing rare genetic diseases using gene panels, clinical exomes, and whole genome sequencing requires rigorous methods of data analysis and interpretation in molecular diagnostics labs. Curating the entire human genome at the gene and variant level will have a significant impact on our understanding of human genetics and how to apply these insights for clinical care. In particular, systematic identification of all published variants and expert review of associated evidence is a critical component in the resolution of variants of uncertain significance (VUS) and will enable more rapid and accurate variant interpretation.
In this talk, Chief Science Officer and Founder, Dr. Mark Kiel will convey how this curation will accelerate the field of genomics, patient diagnosis, and precision medicine development.
Following, Brittnee Jones, VP of Product Management, and Jeffrey Bissonnette, Senior Director of Genomic Curation, will deliver a comprehensive overview of Genomenon’s pioneering approach to curating the entire genome, providing insight into both known and novel variants.
EXHIBIT THEATER SPEAKERS



Mark Kiel, MD, PhD
Founder and Chief Scientific Officer
Jeffrey Bissonnette, MSc, CGC Senior Director of Genomic Curation
Brittnee Jones, PhD
VP of Product Management and Product Strategy
Scientific Platform Presentation
Comprehensive identification of gene-disease relationships across the clinical exome through systematic literature review and parallelized evidence curation
Sequencing of large gene panels, exomes and genomes in clinical diagnostics, has led to an exponential increase in the number of variants of uncertain significance. Additionally, with the commoditization of genome sequencing, newly characterized Gene-Disease Relationships (GDRs) are being published at an exponential pace. Despite the great progress made by multiple expert ClinGen working groups in identifying GDRs from >1900 genes, more work remains to fully characterize newly emerging GDRs from the scientific and medical literature and to stay current with the latest published evidence to keep designations up-to-date. For this study, we embarked on massively parallelized curation of all GDRs across all genes associated with the clinical exome using a gene-first approach facilitated by computational indexing of published evidence to ensure maximal sensitivity.

WATCH NOW | Genomenon’s Scientific Platform Presentation
and Exhibit Theater talks
Posters:
An estimation of global prevalence of PLA2G6-associated neurodegeneration
Friday, March 15, 2024 | 10:30AM – 12:00PM EST
In this study we estimated the genetic prevalence of PLA2G6-associated neurodegeneration, which is an early onset progressive neurodegenerative disease. The overall genetic prevalence was estimated to be 1 in 220,322 pregnancies, with the highest genetic prevalence in African/African-American populations at 1 in 86,012 pregnancies. We found a significant underdiagnosis of PLA2G6-associated neurodegeneration, especially in the non-European populations. Thus, expanded genetic sequencing is needed in non-European populations for accurate diagnosis.
Clarification of variant reporting for homologous genes resolved through systematic literature review – ACMG SF genes CALM1, CALM2, and CALM3
Friday, March 15, 2024 | 10:30AM – 12:00PM EST
We report on the findings of semi-automated curation of the published dataset for the secondary finding (SF) genes CALM1, CALM2, and CALM3 (CALM) associated with severe calmodulinopathies. These recent additions to the ACMG SF list encode an identical calmodulin protein, and only differ in their promoter and untranslated regions. Published journal articles with at least one variant in CALM, as identified by the Mastermind Genomic Intelligence Platform, were evaluated with additional assessment of clinically-encountered variants in ClinVar. Evidence was assessed using AI-driven identification of relevant articles, automated assignment of population and computational-based ACMG categories, and expert review of literature by our curation specialists. The variant list was filtered to remove benign, synonymous, non-coding variants, and/or variants lacking clinical evidence of disease-association with Long QT Syndrome (LQTS) or Catecholaminergic Polymorphic Ventricular Tachycardia (CPVT). Finally, the presence of coding missense variants across each of the CALM genes, and their interpretations, were quantified and analyzed.
As CALM genes encode the same protein with similar cardiac expression one can imply that a P/LP variant in one gene would be causative to another CALM gene. The idea of homologous-annotation is novel to this gene set but not as a concept. Homologous-annotation, the application of pathogenic criteria in variants across homologous genes with related/identical protein-function, is necessary to minimize missed diagnoses.The application of homologous-annotation across CALM genes increased the total P/LP variant count by over 200%. Consistent application of homologous-annotation in curation guidelines is necessary to avoid missed diagnoses for CALM genes associated with sudden death.
Meet us at Booth #911
Connect with members of our leadership team at ACMG 2024! Talk to a Mastermind Specialist and:
Demo the world’s most comprehensive source of genomic evidence, Drop by for a 1:1 or join us for a larger group demo at:
• Thursday, March 14 @ 10:15am
• Thursday, March 14 @ 1:15pm
•Thursday, March 14 @ 3:15pm
• Friday, March 15 @ 10:30am
Schedule 1:1 meetings – on-site or virtually
Learn about extended trials and special offers for clinical labs

ACMG RESOURCES
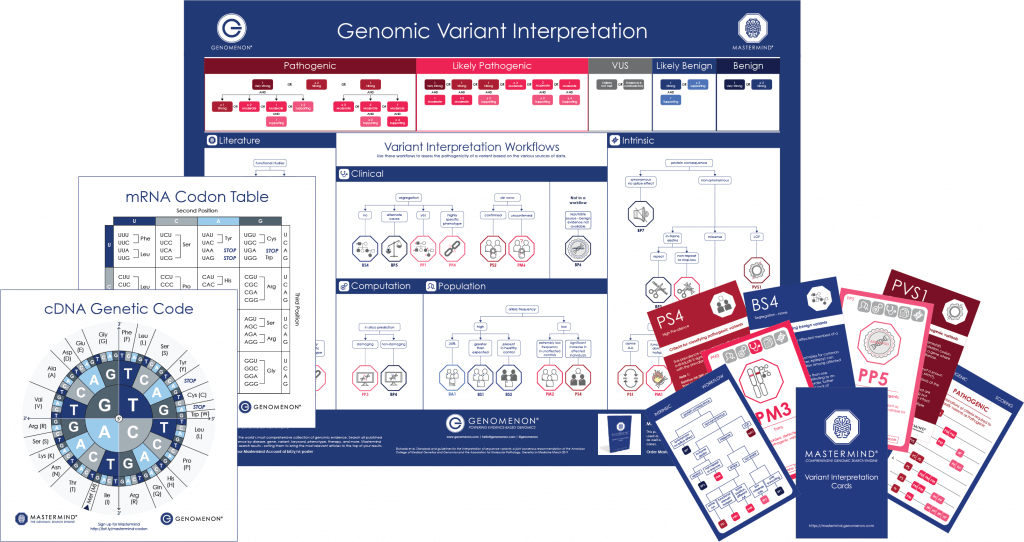
Decorate your workspace with our ACMG Poster, or test your knowledge with our Variant Interpretation Cards.
Even better – get your own copy of our popular cDNA Codon Charts and Intronic Variant Reference Guide when you visit our booth.
Genomenon Networking Reception at ACMG 2024
Thursday, March 14 | 6:00 – 8:00pm EST
Join us for an evening mixer at the Legends Lounge! Situated inside Toronto Marriott City Centre hotel, overlooking Rogers Centre, home of the Toronto Blue Jays, we welcome our valued customers and partners to connect with us. Registration is required and space is limited.
Contact Us for Information

