In an exciting new development, the Mastermind Genomic Search Engine has been featured in DNASTAR’s new eBook, 7 Steps for Human Variant Analysis (and How to Automate Them!)
Written for a diverse range of variant analysts, the instructions and considerations in this eBook are highly relevant for those involved in the process of deriving meaning from genetic data – whether you’re at a core facility, within a bioinformatics group, or by yourself. Specifically, the book explores the challenges involved in genetic variant interpretation, and digs deep into some of the innovative solutions available for streamlining the process – with a special focus on Genomenon’s AI-driven genomic technology found in Mastermind.
Mastermind provides immediate insight into millions of full-text articles encompassing the entire breadth and depth of published genomic evidence. Using patented algorithms, Mastermind allows you to filter and prioritize relevant articles with high specificity, without compromising sensitivity. Additionally, it is able to draw genetic associations between keywords (e.g. genes, variants, diseases, CNVs, phenotypes, and therapies) and deliver a segmented literature output that has been customized for your particular case.
So, with the recent addition of Mastermind to the Lasergene Genomics variant analysis pipeline, the once labor-intensive task of variant interpretation is now faster, easier, and more accurate. In case you missed the news of this recent synergy, you can read the blog from our Mastermind Connections series.
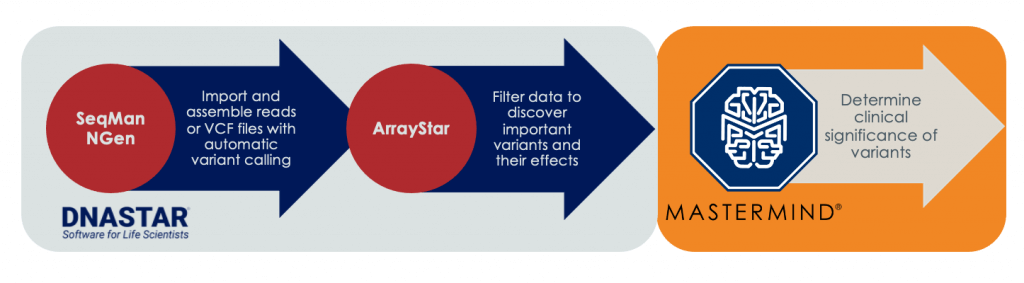
With this in mind, a key highlight of the DNASTAR eBook includes a thorough demonstration of how this workflow integration actually works. As seen below, SeqMan NGen is first used to import and assemble reads or VCF files with automatic variant calling, followed by filtering with ArrayStar. Then, Mastermind uses AI-driven genomics to determine the clinical significance of these variants, as well as draw actionable associations.
Using Mastermind to Unpack a Clinical Case
But how does this process apply to a clinical example? Perhaps the most compelling feature of the eBook is Chapter 3, which demonstrates the power of Mastermind’s integration with a clinical use case. As an example, the authors describe a recent collaboration between DNASTAR and Resonant Therapeutics that used RNA-Seq Illumina reads to study gene expression differences in ovarian carcinoma cells.
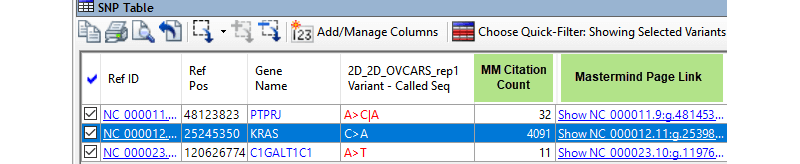
Going through the streamlined 3-step process, the authors report a total of 965,209 variants called using SeqMan NGen, followed by variant filtration in ArrayStar. As seen in the output table below, the G12V variant in the KRAS gene was shown to have 4,000+ Mastermind citations, and is the variant found to be most involved in progression of these particular ovarian tumors.
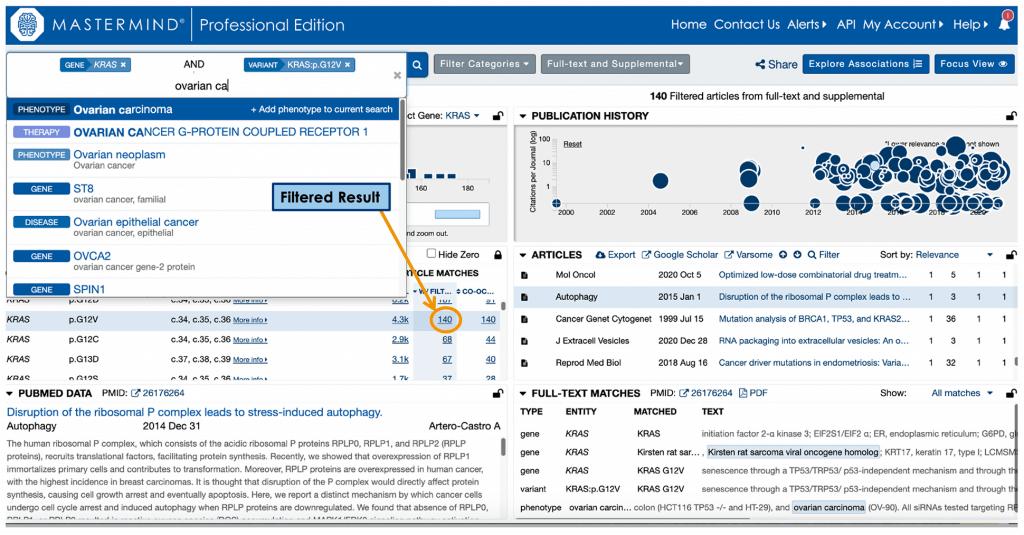
The great thing about this is that ArrayStar includes a direct link to Mastermind, so users can view the entry for known KRAS mutations. Categorical keywords can then be used for further narrowing – for example, as seen below, when the phenotype “ovarian carcinoma” is applied, the reference number is reduced dramatically to 140 relevant articles.
From here, the eBook example shows further refinement of the search criteria to a smaller list of highly relevant articles. Each additional filter applied returns more specific results.
Exploring Ovarian Carcinoma Therapies with Genomic Associations
For clinical decision makers, knowing which therapies are the most effective for a specific mutation can increase precision in patient care. This awareness can be a great starting point for researchers as they begin to form hypotheses in this area.
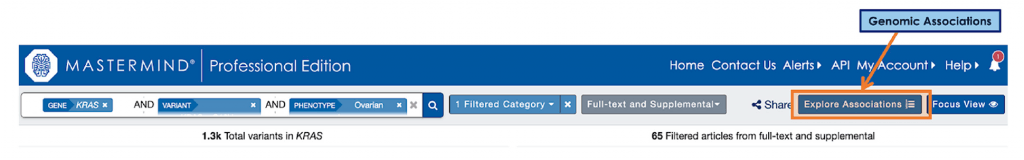
The DNASTAR eBook expands on the clinical case example by introducing this question of therapeutics – in particular, what treatment options are available for ovarian carcinoma related to KRAS G12V mutations? As seen below, by clicking on the blue “Explore Associations” box, the user is directed to a new feature added to Mastermind called Genomic Associations, a flexible user interface that connects search keywords with all associated genetic factors in the medical evidence.
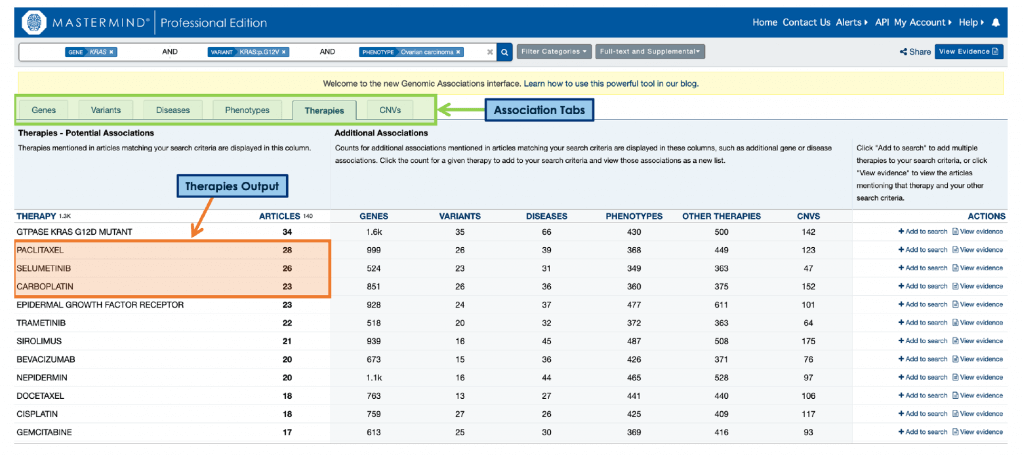
From this example, it is possible to see the top therapies specifically associated with ovarian carcinoma related to KRAS p.G12V mutations (paclitaxel, selumetinib, and carboplatin), as well as explore the associated cited literature. This feature is perfect for anyone interested in generating targeted treatment plans, from genetic researchers to clinical practitioners, and is a powerful addition to DNASTAR’s variant analysis workflow. To learn more about how this interactive “virtual colleague” can help you bring precision to patient care, you can read the blog here.
This is Just the Beginning
Although these patient-facing applications are compelling, they represent a very small part of this technology’s complete potential. As you can see by the other five association tabs, there’s so much more information for Lasergene and Mastermind users to dive into.
To read about these use cases in more detail, as well as learn how you can supercharge your variant interpretation experience, you can download your copy of the eBook directly from DNASTAR. Explore and Discover!